Chromatic Aberration
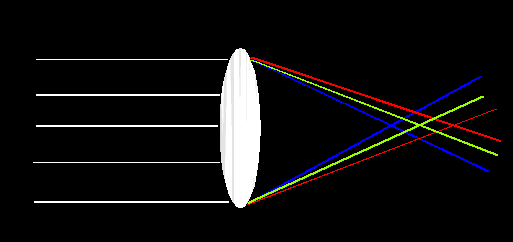
More than 200 years ago, Newton showed that white light was composed of multiple wavelengths. Simple lenses will refract (bend) light differentially as a function of wavelength. Short (blue appearing) wavelengths are refracted more than long (red appearing) wavelengths. Consequently, lenses like the one shown above will not image light all in one place.
A lens called an achromatic lens solves this problem.
You may be interested to know that the human eye lens exhibits chromatic aberration. Fortunately, a yellow pigment in the fovea called the macula lutea helps to protect us from this problem. Yellow pigments absorb blue light.
Color/Color Vision
Table of Contents
Subject Index
Table of Contents [When not using frames]

